* = Undergraduate Student Authors Mentored by Dr. Ay
Journal papers:
Russel, W.*, Perry, J.*, Bonzani, C.*, Dontino, A., Mekonnen, Z., Ay, A. & Taye, B. (2023). Feature Selection and Association rule learning Identify Risk Factors of Malnutrition Among Ethiopian Schoolchildren. Frontiers in Epidemiology, 3: 1150619.
Wang, D.*, Russel, W. A.*, Sun, Y.*, Belanger, K. D., & Ay, A. (2023). Machine learning and network analysis of the gut microbiome from patients with schizophrenia and non-psychiatric subject controls reveal behavioral risk factors and bacterial interactions. Schizophrenia Research, 251, 49-58.
Jimenez, A. G., Paul, K., Zafar, A.*, & Ay, A. (2023). Effect of different masses, ages, and coats on the thermoregulation of dogs before and after exercise across different seasons. Veterinary Research Communications, 47(2), 833-847.
Overton, R., Zafar, A.*, Attia, Z.*, Ay, A., & Ingram, K. K. (2022). Machine Learning Analyses Reveal Circadian Features Predictive of Risk for Sleep Disturbance. Nature and Science of Sleep, 1887-1900.
Dhawka, L., Cha, Y.*, Ay, A., Ingram, K.K. (2022). Low circadian amplitude and delayed phase are linked to Seasonal Affective Disorder (SAD). Journal of Affective Disorders Reports, p.100395.
Tran, V.*, Saad, T.*, Tesfaye, M., Walelign, S., Wordofa, M., Abera, D., Desta, K., Tsegaye, A., Ay, A., & Taye, B. (2022). Helicobacter pylori (H. pylori) risk factor analysis and prevalence prediction: a machine learning-based approach. BMC Infectious Diseases, 22(1), pp.1-14.
Zinani, O. Q., Keseroglu, K., Dey, S., Ay, A., Singh, A., & Ozbudak, E. M. (2022). Gene copy number and negative feedback differentially regulate transcriptional variability of segmentation clock genes. iScience, 25(7), 104579.
Zafar, A.*, Attia, Z.*, Tesfaye, M., Walelign, S., Wordofa, M., Abera, D., Desta, K., Tsegaye, A., Ay, A., & Taye, B. (2022). Machine learning-based risk factor analysis and prevalence prediction of intestinal parasitic infections using epidemiological survey data. PLoS Neglected Tropical Diseases, 16(6), e0010517.
Birenbaum, Z.*, Do, H.*, Horstmyer, L., Orff, H., Ingram, K., & Ay, A. (2022). SEALNET: Facial recognition software for ecological studies of harbor seals. Ecology and Evolution, 12(5), e8851.
Zafar, A.*, Overton, R., Attia, Z.*, Ay, A., & Ingram, K. (2022). Machine learning and expression analyses reveal circadian clock features predictive of anxiety. Scientific Reports, 12(1), 1-11.
Ren, Y., Sarkar, A., Veltri, P., Ay, A., Dobra, A., & Kahveci, T. (2021). Pattern discovery in multilayer networks. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 19(2), 741-752.
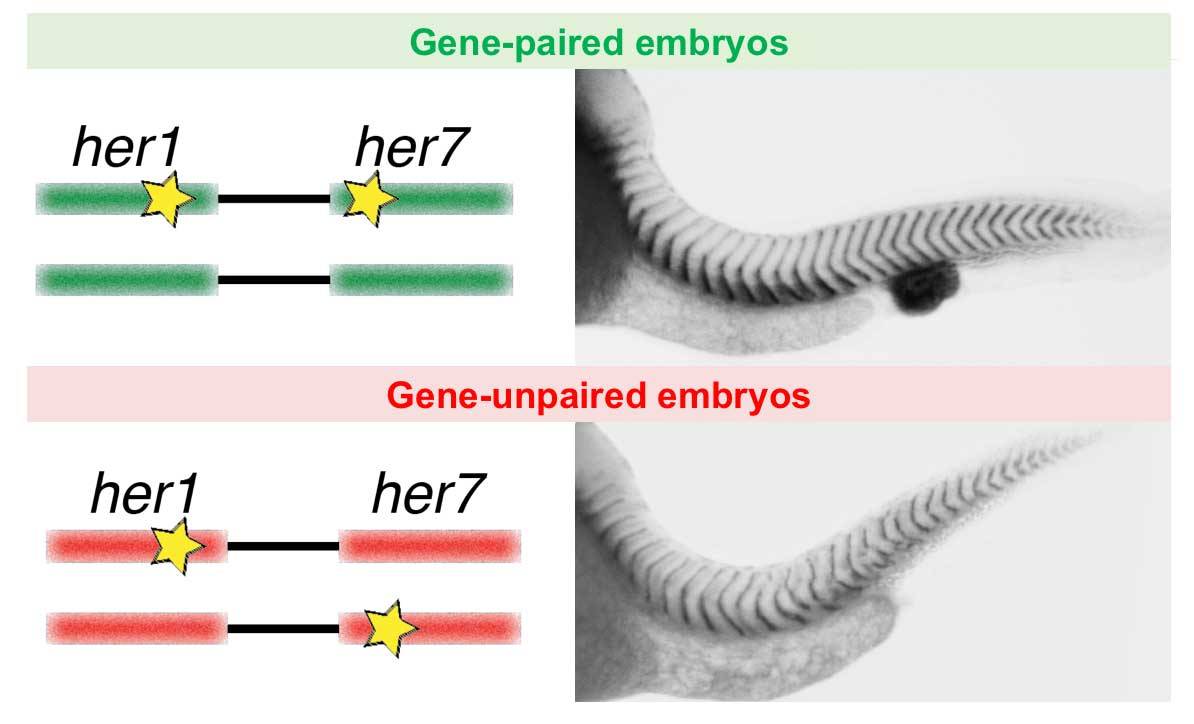
Zinani, O. Q., Keseroğlu, K., Ay, A., & Özbudak, E. M. (2021). Pairing of segmentation clock genes drives robust pattern formation. Nature, 589(7842), 431-436.
Chow, K.*, Sarkar, A., Elhesha, R., Cinaglia, P., Ay, A., & Kahveci, T. (2019). ANCA: Alignment-Based Network Construction Algorithm. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 18(2), 512-524.
Ren, Y., Ay, A., Dobra, A., & Kahveci, T. (2019). Characterizing building blocks of resource constrained biological networks. BMC Bioinformatics, 20(12): 318.
Keskin, S., Simsek, M. F., Vu, H. T.*, Yang, C.*, Devoto, S. H., Ay, A., & Özbudak, E. M. (2019). Regulatory network of the scoliosis-associated genes establishes rostrocaudal patterning of somites in zebrafish. iScience, 12, 247-259.
Ren, Y., Ay, A., & Kahveci, T. (2018). Shortest path counting in probabilistic biological networks. BMC Bioinformatics, 19(1), 465.
Ren, Y., Ay, A., Gerke, T. A., & Kahveci, T. (2018). Identification of jointly correlated gene sets. Journal of Bioinformatics and Computational Biology, 16(05), 1840019.
Keskin, S., Devakanmalai, G. S., Kwon, S. B.*, Vu, H. T.*, Hong, Q., Lee, Y. Y., Soltani, M., Singh, A., Ay, A., & Özbudak, E. M. (2018). Noise in the vertebrate segmentation clock is boosted by time delays but tamed by notch signaling. Cell Reports, 23(7), 2175-2185.
Liberman, A. R.*, Halitjaha, L.*, Ay, A., & Ingram, K. K. (2018). Modeling strengthens molecular link between circadian polymorphisms and major mood disorders. Journal of Biological Rhythms, 33(3), 318-336.
Alim, M. A., Ay, A., Hasan, M. M., Thai, M. T., & Kahveci, T. (2017). Construction of Signaling Pathways with RNAi Data and Multiple Reference Networks. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 15(4), 1079-1091.
Zaifman, J.*, Shan, D.*, Ay, A., & Jimenez, A. G. (2017). Shifts in bird migration timing in North American long-distance and short-distance migrants are associated with climate change. International Journal of Zoology, 6025646.
Liberman, A. R.*, Kwon, S. B.*, Vu, H. T.*, Filipowicz, A., Ay, A., & Ingram, K. K. (2017). Circadian clock model supports molecular link between PER3 and human anxiety. Scientific Reports, 7(1), 1-10.
Armstrong, G. W.*, Mahmood, A.*, Nugent, A., Dexter, S., Hutto, E., McCay, T. S., Ay, A. (2017) WORMSPREAD: an individual-based model of invasive earthworm population dynamics. Computational Ecology and Software 7(3): 109-122.
Yildirim, N., Aktas, M. E., Ozcan, S. N., Akbas, E., & Ay, A. (2017). Differential transcriptional regulation by alternatively designed mechanisms: A mathematical modeling approach. In Silico Biology, 12(3-4), 95-127.
Ren, Y., Wang, Q., Hasan, M. M., Ay, A., & Kahveci, T. (2016). Identifying the topology of signaling networks from partial RNAi data. BMC Systems Biology, 10(2), 231-242.
Ingram, K. K., Ay, A., Kwon, S. B.*, Woods, K., Escobar, S., Gordon, M., Smith I., Bearden I., Filipowicz A., & Jain, K. (2016). Molecular insights into chronotype and time-of-day effects on decision-making. Scientific Reports, 6(1), 1-9.
Belanger, K. D., Larson, N.*, Kahn, J.*, Tkachev, D., & Ay, A. (2016). Mutant screen report: Microarray analysis of gene expression in S. cerevisiae kap108Delta mutants upon addition of oxidative stress. G3: Genes, Genomes, Genetics, 6(4), 1131-1139.
Kok, K., Ay, A., Li, L. M., & Arnosti, D. N. (2015). Genome-wide errant targeting by Hairy. eLife, 4, e06394.
Ferrante, A., Gellerman, D., Ay, A., Woods, K. P., Filipowicz, A. M., Jain, K., Jain K., Bearden N., & Ingram, K. K. (2015). Diurnal preference predicts phase differences in expression of human peripheral circadian clock genes. Journal of Circadian Rhythms, 13(4), 1-7.
Ay, A., Wilner, N.*, & Yildirim, N. (2015). Mathematical modeling deciphers the benefits of alternatively-designed conserved activatory and inhibitory gene circuits. Molecular BioSystems, 11(7), 2017-2030.
Ay, A., Gong, D., & Kahveci, T. (2015). Hierarchical decomposition of dynamically evolving regulatory networks. BMC Bioinformatics, 16(1), 1-19.
Ay, A., Holland, J.*, Sperlea, A.*, Devakanmalai, G. S., Knierer, S., Sangervasi, S.*, Stevenson A., & Özbudak, E. M.(2014). Spatial gradients of protein-level time delays set the pace of the traveling segmentation clock waves. Development, 141(21), 4158-4167.
Ay, A., Gong, D., & Kahveci, T. (2014). Network-based prediction of cancer under genetic storm. Cancer Informatics, 13(Suppl 3), 15–31.
Ay, A., & Yildirim, N. (2014). Dynamics matter: differences and similarities between alternatively designed mechanisms. Molecular BioSystems, 10(7), 1948-1957.
Ay, A., Knierer, S., Sperlea, A.*, Holland, J.*, & Özbudak, E. M. (2013). Short-lived Her proteins drive robust synchronized oscillations in the zebrafish segmentation clock. Development, 140(15), 3244-3253.
Suleimenov, Y.*, Ay, A., Samee, M. A. H., Dresch, J. M., Sinha, S., & Arnosti, D. N. (2013). Global parameter estimation for thermodynamic models of transcriptional regulation. Methods, 62(1), 99-108.
Dresch, J. M., Liu, X.*, Arnosti, D. N., & Ay, A. (2010). Thermodynamic modeling of transcription: sensitivity analysis differentiates biological mechanism from mathematical model-induced effects. BMC Systems Biology, 4(1), 1-11.
Fakhouri, W. D., Ay, A., Sayal, R., Dresch, J., Dayringer, E.*, & Arnosti, D. N. (2010). Deciphering a transcriptional regulatory code: modeling short‐range repression in the Drosophila embryo. Molecular Systems Biology, 6(1), 341.
Ay, A., Fakhouri, W. D., Chiu, C., & Arnosti, D. N. (2008). Image processing and analysis for quantifying gene expression from early Drosophila embryos. Tissue Engineering Part A, 14(9), 1517-1526.
Ay, A., Gürses, M., & Zheltukhin, K. (2003). Hamiltonian equations in R3. Journal of Mathematical Physics, 44(12), 5688-5705.
Conference papers:
Ren, Y., Sarkar, A., Ay, A., Dobra, A., & Kahveci, T. (2019) Finding conserved patterns in multilayer networks. Proceedings of the 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics, pp. 97-102
Chow, K.*, Ay, A., Elhesha, R., & Kahveci, T. (2018) ANCA: Alignment-based Network Construction Algorithm. Proceedings of the 9th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics, pp. 21-26
Ren, Y., Ay, A., Dobra, A. & Kahveci, T. (2018) Characterizing building blocks of resource constrained biological networks. Proceedings of the 9th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics, pp. 581-582
Ren, Y., Ay, A., Gerke, T., & Kahveci T (2018) Searching jointly correlated gene combinations. Proceedings of the 10th International Conference on Bioinformatics and Computational Biology, BICOB.
Wang, Q., Ren, Y., Hasan, M. M., Ay, A., & Kahveci, T. (2015) Construction of signaling networks with incomplete RNAi data. Proceedings of the Bioinformatics and Biomedicine (BIBM) - 2015 IEEE International Conference, pp. 157-162.
Alim, M. A., Ay, A., Hasan, M. M., Thai, M.T., & Kahveci, T. (2015) Multiple reference networks improve accuracy of signaling network construction. Proceedings of the 6th ACM Conference on Bioinformatics - Computational Biology and Health Informatics, pp. 176-185.
Review papers:
Dresch, J. M., Richards, M.*, & Ay, A. (2013). A primer on thermodynamic-based models for deciphering transcriptional regulatory logic. BBA Gene Regulatory Mechanisms, 1829(9), 946-953.
Ay, A., & Arnosti, D. N. (2011). Mathematical modeling of gene expression: a guide for the perplexed biologist. Critical Reviews in Biochemistry and Molecular Biology, 46(2),
Commentaries:
Arnosti, D. N., & Ay, A. (2012) Boolean modeling of gene regulatory networks: Driesch redux. PNAS 109(45), 18239-18240.
Ay, A., & Arnosti, D. N. (2010) Nucleosome positioning: An essential component of the enhancer regulatory code? Current Biology 20(9), R404-406.