“Nothing in biology makes sense except in the light of evolution,” wrote famed geneticist Theodosius Dobzhansky in 1973.
Associate Professor of Biology and Mathematics Ahmet Ay and colleagues at the Cincinnati Children’s Hospital have demonstrated this tenet of biology once again in their recent Nature paper, published last month. In it, they examine a common genetic pattern, called paired genes, that occurs across all organisms. They demonstrate for the first time how this gene pattern can be advantageous, if not critical, to an organism’s development.
The gene pair the team studied, her1 and her7, belong to a group called clock genes, so named because negative feedback loops cause their expression to oscillate in cycles. That means, as the genes are expressed — that is, the DNA is transcribed into mRNA, a messenger molecule, which will then be translated into proteins, the building blocks of the cell — the output signals back to the cell that it should slow down expression of those genes. As expression slows and the output lessens, the dampening signal also slows, allowing expression to ramp back up again.
The most famous clock genes are related to circadian rhythms, which cycle in a 24-hour (it’s actually 24.2 hours, says Ay) period and affect human states like sleep and hunger. But in the recent study, Ay and his team studied segmentation clock genes that are important for development, specifically for the growth of consecutive body segments in embryos, using zebrafish as their model organism. In zebrafish, the her1 and her7 genes cycle in 30-minute periods, telling the developing fish’s cells when to make new segments called somites (think: vertebrae).
The interesting aspect of her1 and her7 isn’t actually the zebrafish embryos or the so-called segmentation clock, explains Ay. It’s how the two genes are paired: They’re found near each other on the same chromosome, and they share the same on/off switch. But why are her1 and her7 — or any other gene pairs found across genomes, for that matter — linked in this way? “You see gene pairing in all different organisms, and in all different systems,” says Ay. “What are the benefits of that?”
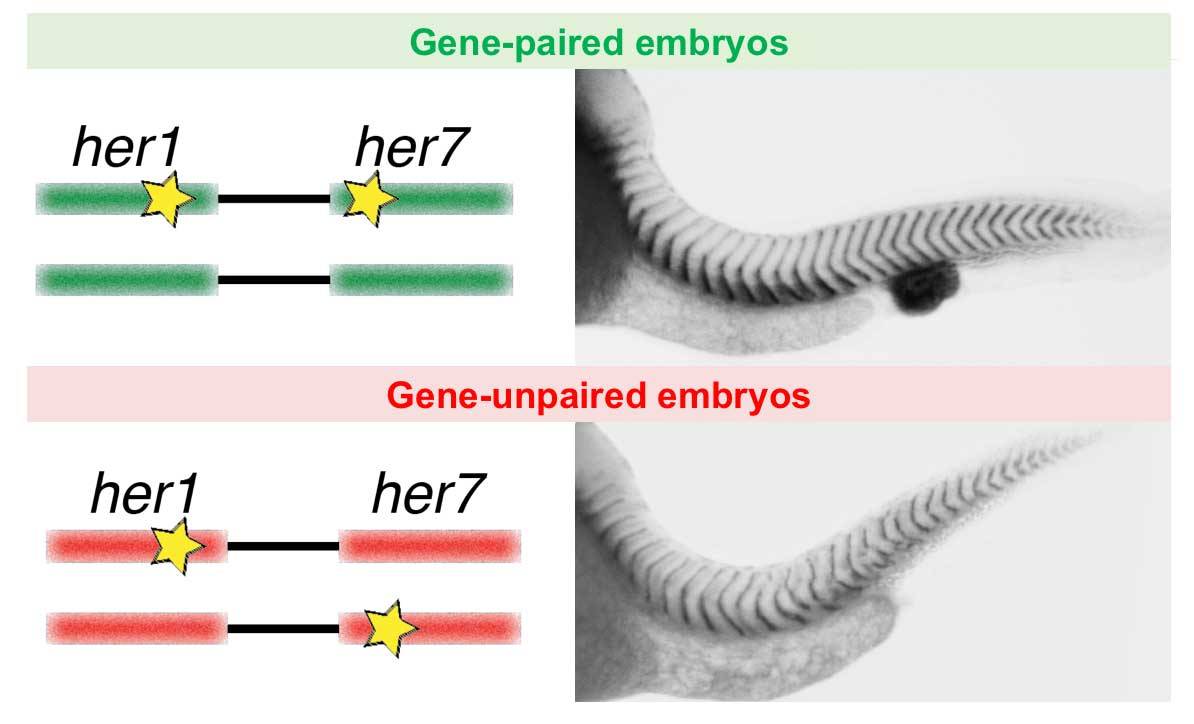
Ay and his collaborators, Oriana Zinani, Kemal Keseroğlu, and Ertuğrul Özbudak, teamed up to figure out why it might be adaptive for these clock genes to be paired. They ran experiments to see what happened when her1 and her7 were unpaired in the cell. Because zebrafish have two copies of each gene in each cell, they could knock out a her1 on one chromosome and a her7 on the other chromosome to see how the cell would respond to having the genes physically separated.
Compared with cells that had a her1 and a her7 paired on the same chromosome, cells with the two genes physically separated couldn’t maintain their oscillations — the clock broke. That’s because when gene expression starts at exactly the same time, it gets a head start at mRNA production that allows it to overshoot the “slow down!” signal. That overshooting is needed to create the oscillations; without it, expression of genes just levels out into a steady state. In other words, the precise simultaneous expression of the two genes together was needed to trigger the oscillations, and this just didn’t work when the genes were separated.
This precision within a single cell also allows these mechanisms to stay precise across all the cells in a growing tissue — which is important for cells in a developing embryo that need to stay in sync.
“[These are] key mechanisms conserved everywhere, so understanding how they work really helps us to understand not one system only, but many different biological systems,” says Ay, whose specialization is in systems biology, mathematical modeling, and statistical learning, which allows his lab group to tackle a wide range of biological questions.
“The thing that draws me to this research is its potential beyond this system, beyond just the zebrafish segmentation clock. These findings could be generalizable to other clocks, other biological systems,” he says.